My Work.
|
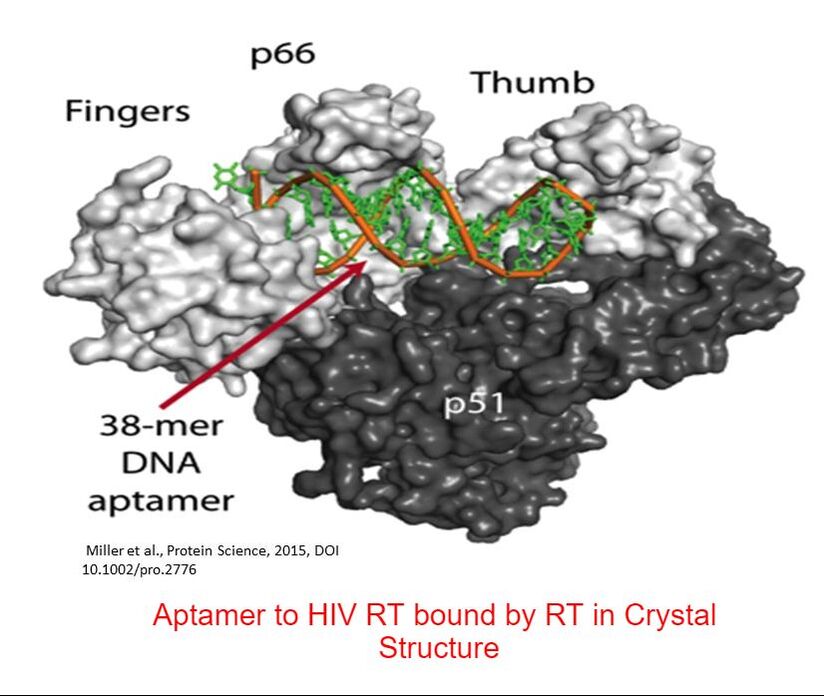
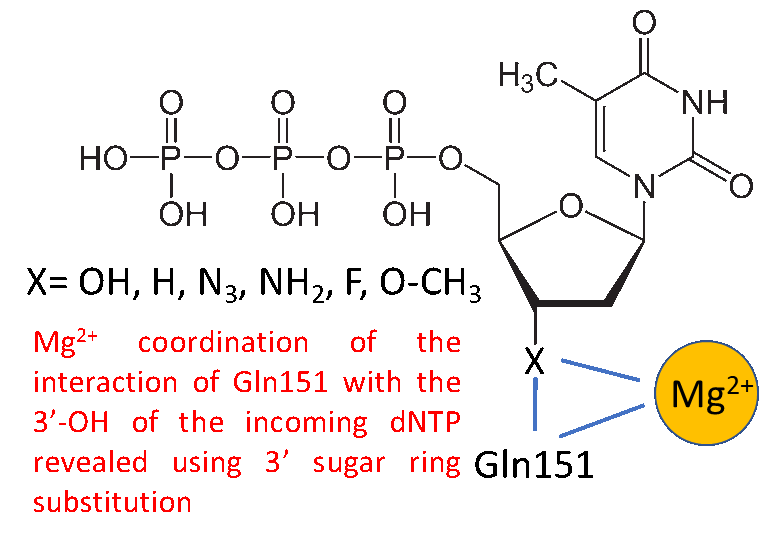
Research in my laboratory has focused on studying the role of HIV-reverse transcriptase (RT) and nucleocapsid protein (NC) in the processes of retroviral recombination and replication. We also analyze basic properties of these proteins including how they interact with specific nucleic acid sequences involved in replication and how the activities of the proteins as defined in the test tube, function in cellular replication. Other projects are aimed at isolating nucleic acid inhibitors/diagnostics (aptamers) that can bind very tightly to viral proteins. potentially being used as diagnostics or inhibitors. We also study poliovirus replication, specifically by examining the polymerase (3Dpol) and 3AB proteins. Viruses in general, and notably HIV, generate high levels of genetic diversity. By producing genetic variants viruses are able to circumvent the host immune response and escape drug therapies. This process is fueled by mutations, many of which are made by the viral polymerase, or RT in the case of HIV. Our lab has focused on understanding how RT creates mutations at a mechanistic level and to what extent RT mutations contribute to the pool of mutations observed during HIV replication. In the last decade, the lab has become increasingly focused on generating unique aptamers. In collaboration with Dr. Phil Holliger (Cambridge University) we were the first to generate fluoroarabino nucleic acid (FANA) aptamers to proteins. FANA is a Xeno-nucleic acid (XNA), that contains a fluorine at the 2' position and an arabinose sugar as opposed to the normal ribose configuration for "normal" nucleotides. XNAs have unique properties that often allow stronger binding to targets and greater resistance to degradation by nucleases in comparison to natural nucleic acids. We are continuing to focus on generating XNA aptamers to viral proteins, including SARS-2 (causative agent of COVID-19). Our goal is to produce XNA aptamers that can be used to study viral proteins, detect viruses in nature, or inhibit viral processes.
|